Regular vs Bayes NN
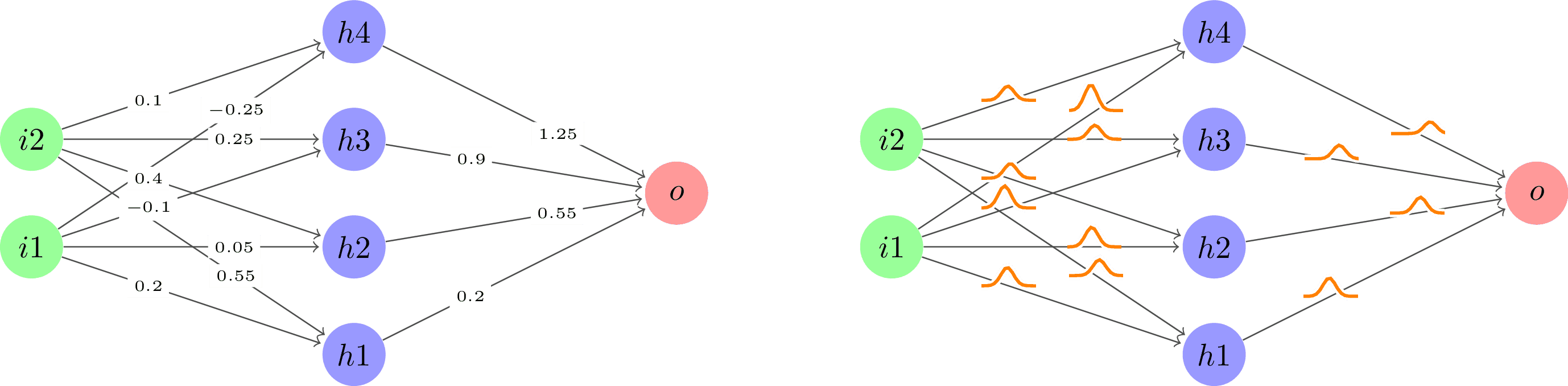
Comparison between regular neural networks and Bayesian neural networks. The diagram illustrates how Bayesian networks incorporate uncertainty in their predictions by treating weights as probability distributions rather than point values, leading to more robust predictions with uncertainty estimates.
Download
Code
regular-vs-bayes-nn.typ (195 lines)
regular-vs-bayes-nn.tex (62 lines)